The ability of a multicellular organism to defend itself against invasion by pathogens (bacteria, fungi, viruses, etc.) depends on its ability to mount immune responses. All metazoans (probably) have inborn defense mechanisms that constitute innate immunity. Vertebrates have not only innate immunity but also are able to mount defense mechanisms that constitute adaptive immunity. [Link]
The hallmarks of adaptive immunity areNot only metazoans, but plants, fungi, and even bacteria have innate mechanisms to defend themselves against pathogens; that is, systems of innate immunity.
Until recently, however, adaptive immunity was thought to be found only in the jawed vertebrates.
But it now turns out that many bacteria and archaea have a mechanism of adaptive immunity. (Both bacteria and archaea lack the membrane-enclosed organelles of eukaryotes and are often called prokaryotes although they are not close relatives).
Bacteria have a number of innate defense mechanisms, for example, the restriction enzymes with which they can destroy foreign DNA that enters their cell without damaging their own DNA. But these defenses show neither great specificity nor memory.
But some bacteria, like E. coli, have a defense system that responds to foreign DNA entering the cell from a virus (bacteriophage) [View] or plasmid and that retains a memory of that encounter through subsequent generations.
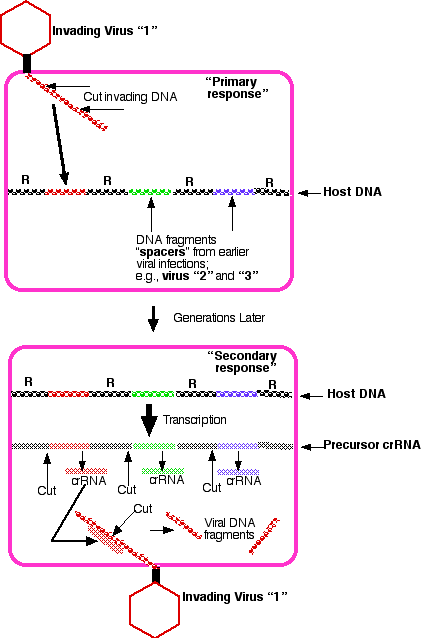
Encoded in the single DNA molecule that is the E. coli genome is a cluster of genetic elements called CRISPR (for clustered regularly interspaced short palindromic repeats). [Link to a discussion of palindromes.] These are indicated by R in the figure.
Between the repeats are so-called spacers that turn out to be segments of DNA that were acquired in the past from an invading virus or plasmid. As the virus inserts its DNA into the cell, enzymes encoded by host genes cut — thus destroying — the invading DNA and then insert one or more pieces between pairs of repeats at one end of the CRISPR loci.
As E. coli continues to divide, its newly-altered CRISPR loci get passed on to subsequent generations.
Transcription of these loci produce long molecules of precursor CRISPR RNA (Precursor crRNA). Host enzymes cut just upstream of each spacer producing a collection of small (~60 nucleotides) CRISPR RNA (crRNA) molecules.
If a cell becomes the target of another virus of the same type ("1"), the crRNA specific for that viral DNA binds to it and, with the help of other enzymes, cuts it into inactive fragments.
Bacteria are under continually assault by viruses and you might expect that over time the CRISPR loci would grow ever-longer. In fact, the older spacers get dropped at about the same rate that new spacers are added.
This RNA-mediated destruction of DNA carrying a complementary sequence shares some parallels with RNA interference by small (or short) interfering RNAs (siRNAs) in eukaryotes [Link] although crRNAs are 2–3 times longer than siRNAs.
The ability of CRISPR RNA to recognize a specific sequence of DNA makes them promising candidates for creating artificial restriction enzymes. Coupled to a DNA-cleaving endonuclease, called Cas9, these molecules can be used to alter specific sites in the genome — adding or deleting or repressing or activating genes. Genome editing with the CRISPR-Cas9 system has worked successfully in a variety of animals, plants and microbes.
| The process and implications of editing eukaryotic genes with the CRISPR-Cas9 system is described in a separate page. Link to it. |
| Read about two other techniques for gene editing: |
| Welcome&Next Search |